Causal variant prioritization for autoimmune target discovery.
InsulatorLeak ranks variants by predicted CTCF insulator disruption—integrating fine-mapping, Enformer, locus-null calibration, and multi-omic validation—so committees get audit-ready, mechanism-ranked outputs.
Show full headline statistics and evidence
>4M participants across 23 GWAS cohorts spanning 7 autoimmune diseases. Fine-mapping, Enformer CTCF SAD, locus null calibration. 45,000× variant reduction. Multi-omic validation. Two independent CRISPR confirmations. 8+ active clinical-stage drug programs across RA, MS, IBD, SLE, T1D, and psoriasis align to InsulatorLeak loci — $25B+ in pharma deals. Patent filed.
Company snapshot
Mechanism-first variant prioritization for autoimmune genetics—packaged as audit-ready scorecards and analyses for target review. Pilot-led go-to-market with a path to broader platform use.
- What we’re building: InsulatorLeak + scoring layers (DQ Score, MPI, CDDI) to rank and explain non-coding signal with explicit mechanistic hypotheses.
- Scale signal: >4M participants across 23 GWAS cohorts (full detail in the Science section below).
- Scientific credibility: Preprint available; details on request.
- Protection / differentiation: patent filed (USPTO) + mechanism-first positioning vs annotation-overlap-first workflows.
- Commercial motion: pilot engagements with credit toward subscription (details under Pharma).
Pharma independently arrived at the same genes — from opposite directions.
InsulatorLeak predicts CTCF insulator disruption from DNA sequence alone. Major pharma programs targeting the same loci were built through biochemical target identification. Zero shared inputs. The convergence is post-hoc and specific — at the level of the exact gene, not the pathway.
RA — PADI4
GAIM p=0.006 from sequence. AstraZeneca’s AZD1163 (anti-PAD2/PAD4 bispecific) is in Phase II targeting PAD4, the enzyme PADI4 encodes.
MS — CD40 (PP.H4=0.958)
InsulatorLeak’s cognate pQTL. Sanofi’s frexalimab (anti-CD40L) is in Phase 3 for MS with data published in NEJM February 2024 — 89% reduction in brain lesions.
IBD — TNFSF15 / IL7R
TNFSF15/TL1A: $21B+ in pharma acquisitions (Roche $7.1B, Merck $10.8B, Sanofi $1.5B). IL7R (34× enrichment): lusvertikimab Phase 2 UC positive data at ECCO 2025.
T1D — IL2RA (PP.H4=0.79)
Cognate pQTL: variant directly controls CD25 protein. Teplizumab (anti-CD3, Sanofi) is FDA approved for Stage 2 T1D — targeting the same IL-2/CD25 T cell axis.
Psoriasis — TYK2 (4 ancestries)
FDR q=0.040 in South Asian MS; replicates 3/4 ancestry groups. Deucravacitinib (BMS, approved PsA). Zasocitinib (Takeda) in Phase 3 for psoriasis and IBD.
SLE — ITGAM (GAIM native)
ONT01 (CD11b/ITGAM agonist, Ontegimod) in Phase 1b/II for lupus nephritis — first-in-class drug at the exact gene InsulatorLeak flagged as a SLE-native insulator locus.
Mechanism-first prioritization: insulator disruption as causal filter.
Annotation-overlap tools ask which SNPs colocalize with regulatory features. We ask which variants causally disrupt insulator function. SuSiE fine-mapping, Enformer CTCF SAD, locus-matched null calibration, chromatin enrichment, eQTL/pQTL colocalization — integrated in one auditable pipeline with per-variant scorecards.
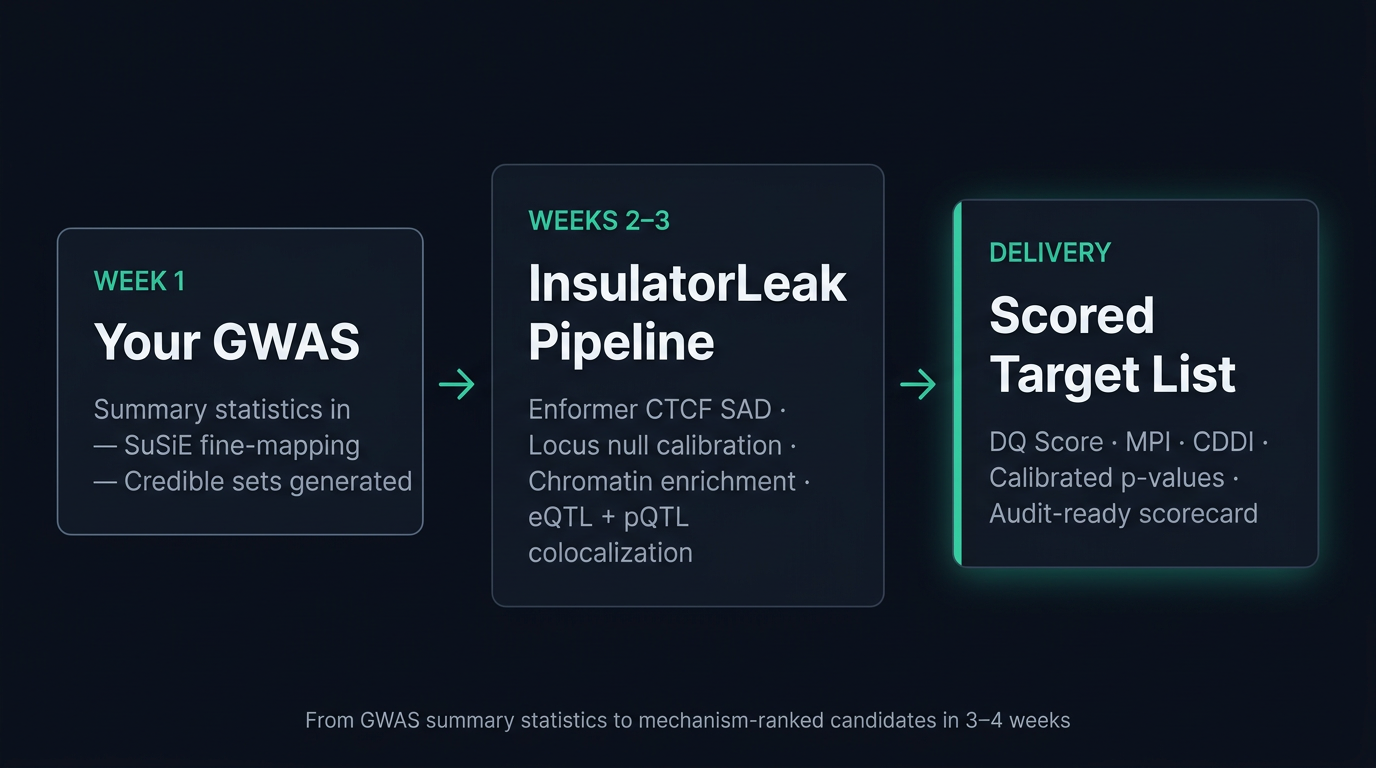
Independent validation layer for target selection.
Locus-matched null calibration distinguishes signal from LD artifact. Multi-cohort replication and cognate pQTL provide variant-to-protein evidence. Output: audit-ready scorecards with calibrated p-values, enrichment, and colocalization — supports go/no-go decisions at committee level.
| Pilot | Target validation package | Enterprise / multi-locus | |
|---|---|---|---|
| Scope | 1–3 loci (disease context agreed upfront) | Same, with deeper per-locus dossier | Multiple loci / recurring cadence |
| Timeline | 4–8 weeks | 4–10 weeks | Quarterly or custom roadmap |
| Deliverables | Per-locus scorecard: replication; Enformer; enrichment; eQTL/pQTL | Above + committee-ready summary + variant tables | Above + integration plan (inputs/outputs, cadence) |
| Decision output | Go / deprioritize / expand | Same + optional follow-up experiments | Portfolio-level view (DQ / MPI / CDDI as applicable) |
| Data / IP | Summary-level inputs; details under CDA | Same | MSA path as you scale |
| Commercial | Pilot fee + credit toward subscription | Package pricing on request | Enterprise subscription on request |
For pharma
Target-validation reports: 1–3 loci, 4–8 weeks. Per-locus scorecard with replication, Enformer, enrichment, eQTL/pQTL. Decision output: go, deprioritize, or expand. Pilot-led; credit toward subscription.
For investors
Mechanism-calibrated variant prioritization at scale. 45,000× reduction. Multi-ancestry, cross-disease validation across MS, IBD, RA, T1D, SLE, psoriasis, and atopic dermatitis. Cognate pQTL validation. Patent filed. Pilot-led GTM with path to platform subscription.
For technical talent
Computational genetics, functional genomics, causal inference. Fine-mapping, regulatory prediction (Enformer, Sei), locus null calibration, multi-omic validation. Pipeline that changes target selection.
Locus null calibration. Multi-omic validation.
Insulator disruption (CTCF SAD) as mechanistic hypothesis; locus-matched nulls for empirical calibration. Chromatin enrichment (Hi-C, ChromHMM) and eQTL/pQTL colocalization for empirical validation. Pan-autoimmune analysis across 7 diseases surfaces shared vs divergent biology.
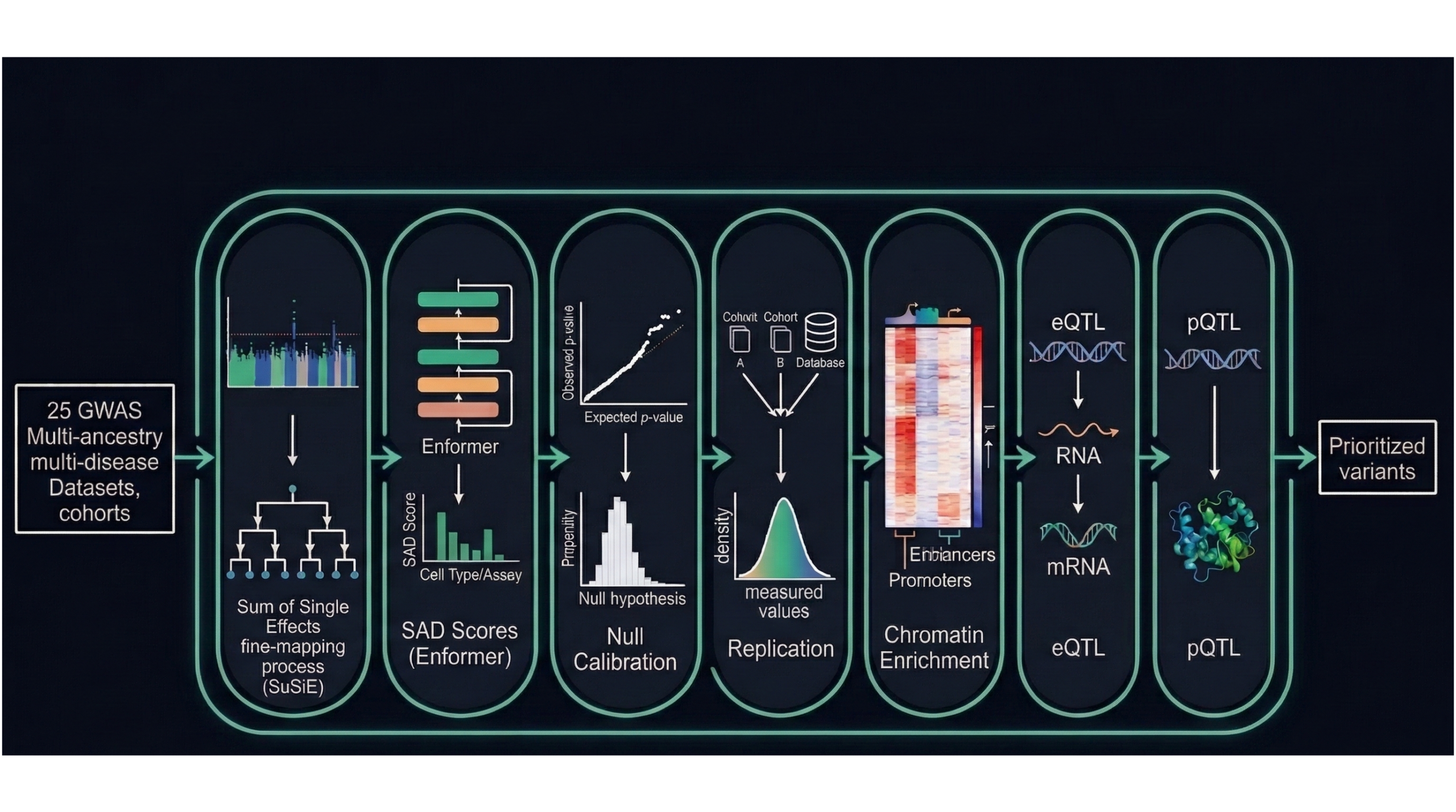
DQ Score
Weighted composite: replication + Enformer + enrichment + eQTL + pQTL. Rule-based, transparent. Audit-ready.
MPI
Mechanism Portfolio Index: dominant mechanism signature per locus. Insulator-loss today; extensible to enhancer/TF.
CDDI
Cross-Disease Divergence Index: MS vs IBD gene prioritization. Same vs different gene — repurposing signal.
Engaging with world-leading genetics centers on validation.
Pipeline output feeds into wet-lab validation: reporter assays, EMSA, allele-specific regulatory activity. Engaging with top-tier genetics and immunology labs on collaborative validation and translational pathways.
Explore collaborationReach out to learn how we can power your drug target discovery.
Whether you’re an investor, a pharma partner, or a technical expert interested in joining — we’d like to hear from you.